
David MacMillan
Contact:
David Drapcho
James S. McDonnell Distinguished University, Professor of Chemistry
[email protected]
Frick Laboratory, 192
609-258-3916
Faculty Assistant:
Jemeille Ackourey
[email protected]
Frick Laboratory, 257
609-258-5152
Research Focus
Research in the MacMillan Lab has evolved over the past few years to encompass bold new investigations in chemical biology that sit alongside our longstanding mission to develop innovative synthetic organic methodologies. The questions that drive our program today confront global challenges for real-world impact, with platforms providing biological insights into cancer, neurodegeneration and aging; new types of reactivity that transform the way we perform chemical reactions and are grounded in molecules that are broadly available; and transformations used throughout the pharmaceutical industry, the agrochemical industry, and in medicinal chemistry. A new partnership with the Ludwig Cancer Research Institute and collaborations with major pharmaceutical companies underscore the scope of our research enterprise.
Areas of active research include:
Biological Insights Using Chemical Biology: Development of µMap
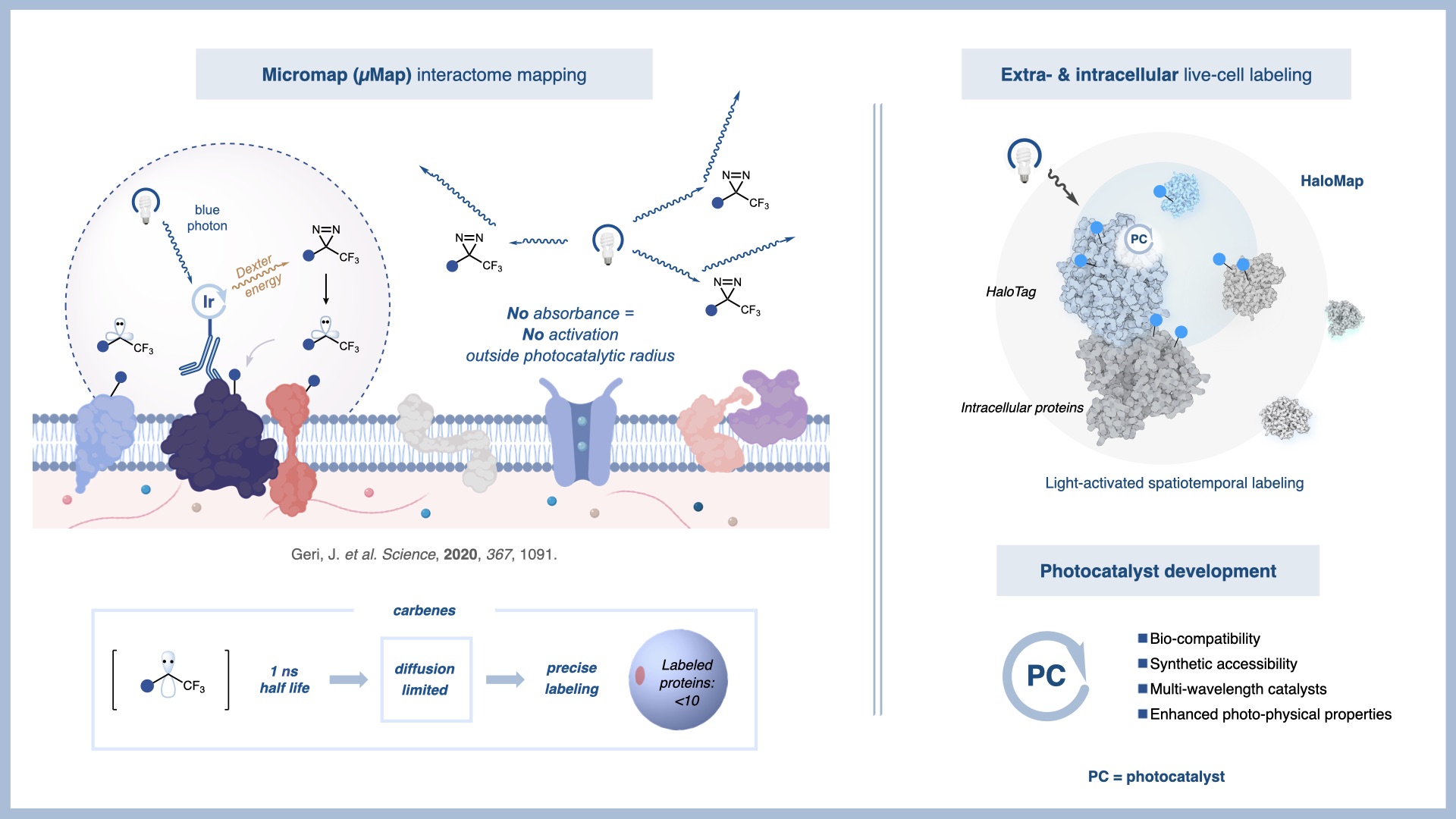
A current focus of the MacMillan group is the development of new chemical tools to probe complex biological systems. In a 2020 Science publication, we introduced a photocatalytic proximity labeling platform, termed µMap, capable of mapping protein–protein interactions in biological systems with nanometer-scale resolution.
Over the past six years, we have expanded µMap into a versatile platform for interrogating biological function. By enabling direct mapping of protein–protein and protein–small molecule interactions in native environments, µMap provides a powerful approach for studying how cells interact with their microenvironment. This technology is now being applied broadly in cancer biology through our collaborations within the Ludwig Princeton Branch.
Ongoing efforts in our laboratory focus on expanding the µMap platform to complex biological settings and applying it to challenging biological problems. Current applications include chromatin biology, intracellular organization, viral host interactions, neuroscience, cancer resistance mechanisms, and cellular stress response. Together, these studies establish µMap as a powerful platform for high-resolution biological discovery. These efforts are supported by extensive collaborations with pharmaceutical and biotechnology partners.
Photoredox Catalysis
Photoredox catalysis has emerged as a powerful platform for chemical synthesis, enabling the use of visible light to generate reactive intermediates under mild conditions. By selectively delivering visible light energy to a photocatalyst, this approach allows facile access to free radicals from abundant feedstock functionalities such as alcohols, olefins, carboxylic acids, and C–H bonds. The versatility of photoredox catalysis has enabled its seamless integration with other powerful activation modes, including organocatalysis, HAT, and transition metal catalysis. As a result, photoredox catalysis is now a central strategy for inventing new chemical transformations that are conceptually novel and broadly applicable across disciplines, ranging from medicinal chemistry to materials science and chemical biology.
Our laboratory has been at the forefront of the development of photoredox catalysis for over 20 years, contributing more than 100 publications to the field, including over 30 papers in Science and Nature.
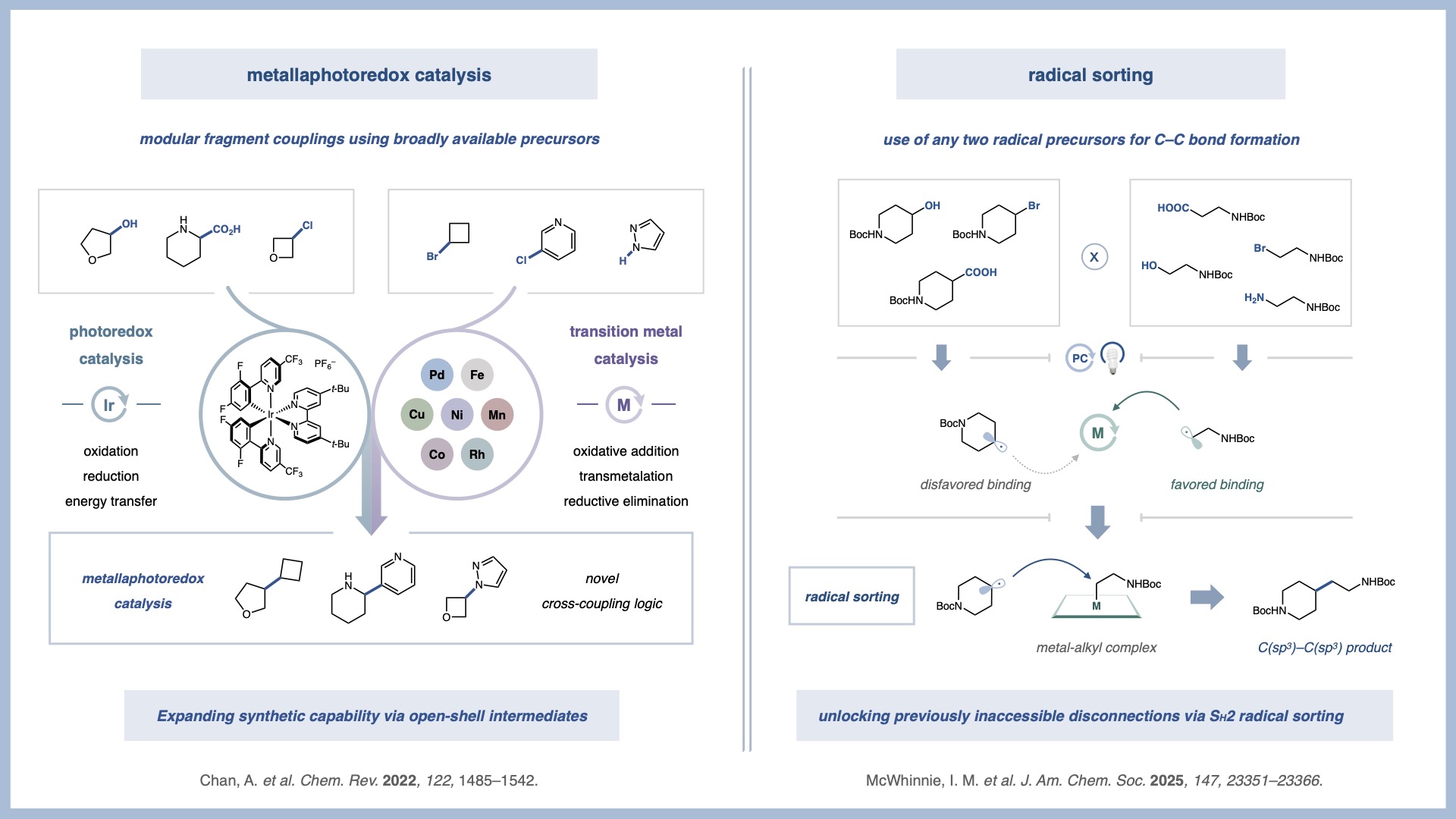
Metallaphotoredox Catalysis
In a 2014 collaboration with the Doyle group, we reported one of the first examples of merging transition metal catalysis and photoredox catalysis to access novel reactivity that cannot be achieved by either catalytic system alone. This new field has delivered a host of highly enabling C–C and C–X bond-forming reactions that reveal previously elusive synthetic disconnections.
Our group’s research in metallaphotoredox catalysis is primarily focused on two metals: nickel and copper. By combining the fundamental steps of nickel catalysis, such as oxidative addition and reductive elimination, with high-energy radicals generated through photoredox catalysis, we have developed novel C(sp2)–C(sp3) and C(sp3)–C(sp3) bond forming reactions. We have also merged copper catalysis with photoredox catalysis, establishing a new catalytic platform with orthogonal reactivity to previous metallaphotoredox systems. The ability to access elusive high-energy copper intermediates under exceedingly mild conditions has unlocked new transformations, including C–CF3, C–X, C–N, and C–C bond formations. In addition, recent efforts have begun to explore iron-based metallaphotoredox catalysis, further expanding the scope of accessible reactivity using earth-abundant metals.
Radical Sorting
As an outgrowth of our metallaphotoredox program, in 2021 we disclosed one of the first applications of SH2 radical sorting to chemical methodology. While SH2 is known in biological systems, it had not yet been harnessed as a general platform for cross-coupling reactions in the laboratory. We showed that this platform could be used to enable previously inaccessible disconnections.
Unlike traditional inner-sphere mechanisms that rely on unstable dialkyl metal complexes, outer-sphere SH2 radical sorting enables the formation of challenging motifs such as all-aliphatic quaternary carbons. To date, we have applied radical sorting to directly merge abundant feedstock functionalities, achieving alcohol–carboxylic acid and alcohol–alcohol cross-coupling. More recently, we have extended this logic to multi-component transformations, including triple radical sorting processes that enable alkene difunctionalization reactions.
Merck Center for Catalysis
To accelerate reaction discovery and optimization, we leverage the Merck Center for Catalysis at Princeton University, a state-of-the-art facility that enables the high-throughput execution and analysis of catalytic reactions. The center houses a Chemspeed Accelerator robotic platform, an automated system used to apply the center’s resources to accelerate challenging, high-value projects for both methodology development and optimization of key steps in total synthesis.
Honors
2021 Nobel Prize in Chemistry
Honorary Fellowship, Royal Society of Chemistry (2026)
Extraordinarius Award, UC Irvine (2024)
F. A. Cotton Medal for Excellence in Chemical Research, ACS (2024)
Chirality Medal Award (2022)
Knight Bachelor (2022)
Centenary Prize, Royal Society of Chemistry, UK (2020)
Nagoya Medal, Nagoya University (2019)
Elected to the National Academy of Sciences, USA (2019)
ACS Gabor Somorjai Award in Catalysis (2018)
Noyori Prize, Japanese Society of Synthetic Chemistry (2018)
Henry J. Albert Award from BASF and IMPI (2017)
Janssen Pharmaceutical Prize, Belgium (2016)
Ohio State Edward Mack Jr. Award (2016)
Tischler Award, Harvard University, MA 2016 ACS Kosolapoff Award (2016)
Hamilton Award in Molecular Sciences, University of Nebraska (2015)
Schering Foundation Prize (Berlin) for Outstanding Research in Medicine, Biology or Chemistry (2015)
NJ ACS Award for Creativity in Molecular Design and Synthesis (2014)
Harrison Howe ACS Award in Chemistry (2013)
Elected to the Fellowship of the Royal Society (FRS, 2012)
Elected to the American Academy of Arts and Sciences (2012)
ACS Prize for Creative Work in Organic Synthesis (2011)
Mitsui Award in Catalysis (2011)
Mukaiyama Prize (Japanese Society of Organic Chemists, 2007)
Arthur C. Cope Scholar ACS Award (2007)
Thieme-IUPAC Prize in Synthetic Organic Chemistry (2006)
Elias J. Corey ACS Award (2005)
Selected Publications
“Couple-Close: Unified Approach to Semi-Saturated Cyclic Scaffolds.” Xie, J.; Zhao, W. Y.; Wang, J. Z.; Lyon, W. L.; Takanashi, N.; Long, A.; Sodano, T. M.; Kelly, C. B.; Bryan, M. C.; MacMillan, D. W. C. Science 2026, 391, 399.
“µMap-FFPE: A High-Resolution Protein Proximity Labeling Platform for Formalin-Fixed Paraffin-Embedded Tissue Samples.” Bissonnette, N. B.; Zamanis, M. E.; Knutson, S. D.; Boyer, Z.; Harris, A.; Martin, D.; Geri, J. B.; Couto, S.; Ahmadi, T.; Muthuswamy, A.; Fereshteh, M.; MacMillan, D. W. C. J. Am. Chem. Soc. 2025, 147, 23387.
“Generalizing arene C−H alkylations by radical-radical cross coupling.” Großkopf, J.; Gopatta, C.; Martin, R. T.; Haseloer, A.; MacMillan, D. W. C. Nature 2025, 641, 112.
“µMap proximity labeling in living cells reveals stress granule disassembly mechanisms.” Pan, C.; Knutson, S. D.; Huth, S. W.; MacMillan, D. W. C. Nat. Chem. Biol. 2024, 21, 490.
“Unlocking Carbene Reactivity by Metallaphotoredox a-Elimination.” Boyle, B. T.; Dow, N. W.; Kelly, C. B.; Bryan, M. C.; MacMillan, D. W. C. Nature 2024, 631, 789.
“Alcohol–alcohol cross-coupling enabled by SH2 radical sorting.” Chen, R.; Intermaggio, N. E.; Xie, J.; Rossi-Ashton, J. A.; Gould, C. A.; Martin, R. T.; Alcázar, J.; MacMillan, D. W. C. Science 2024, 383, 1350.
“Alkene dialkylation by triple radical sorting.” Wang, J. Z.; Lyon, W. L.; MacMillan, D. W. C. Nature 2024, 628, 104
“Exploiting the Marcus inverted region for first-row transition metal-based photoredox catalysis.” Chan, A. Y.; Ghosh, A.; Yarranton, J. T.; Twilton, J.; Jin, J.; Arias-Rotondo, D. M.; Sakai, H. A.; McCusker, J. K.; MacMillan, D. W. C. Science 2023, 382, 191.
“Tracking Chromatin State Changes Using Nanoscale Photo-Proximity Labelling.” Seath, C. P.; Burton, A. J.; Sun, X.; Lee, G.; Kleiner, R. E.; MacMillan, D. W. C.; Muir, T. W. Nature 2023, 616, 574.
“Rapid and Modular Access to Quaternary Carbons from Tertiary Alcohols via Bimolecular Homolytic Substitution.” Gould, C. A.; Pace, A. L.; MacMillan, D. W. C. Am. Chem. Soc. 2023, 145, 16330.
“General Access to Cubanes as Benzene Bioisosteres.” Wiesenfeldt, M. P.; Rossi-Ashton, J. A.; Perry, I. B.; Diesel, J.; Garry, O. L.; Bartels, F.; Coote, S. C.; Ma, X.; Yeung, C. S.; Bennett, D. J.; MacMillan, D. W. C. Nature 2023, 618, 513.
“µMap Photoproximity Labeling Enables Small Molecule Binding Site Mapping.” Huth, S. W.; Oakley, J. V.; Seath, C. P.; Geri, J. B.; Trowbridge, A. D.; Parker, D. L.; Rodriguez-Rivera, F. P.; Schwaid, A. G.; Ramil, C.; Ryu, K. A.; White, C. H.; Fadeyi, O. O.; Oslund, R. C.; MacMillan, D. W. C. Am. Chem. Soc. 2023, 145, 16289.
“Metallaphotoredox-enabled deoxygenative arylation of alcohols” Dong, Z.; D. W. C. MacMillan; Nature 2021, 598, 451.
“Microenvironment mapping via Dexter energy transfer on immune cells.” Geri, J. B.; Oakley, J. V.; Reyes-Robles, T; Wang, T.; McCarver, S.; White, C. H.; Rodriguez-Rivera, F. P.; Parker D. L. Jr.; Hett, E. C.; Fadeyi, O. O.; Oslund, R. C. W. C. MacMillan; Science 2020, 367, 1091.
“A Radical Approach to the Copper Oxidative Addition Problem: Trifluoromethylation of Bromoarenes.” Le, C.; Chen, T. Q.; Liang, T.; Zhang, P.; MacMillan, D. W. C. Science 2018, 360, 1010.
“Direct Arylation of Strong Aliphatic C–H Bonds.” Perry, I. B.; Brewer, T. F.; Sarver, P. J.; Schultz, D. M.; DiRocco, D. A.; MacMillan, D. W. C. Nature 2018, 560, 70.
“Photosensitized, energy transfer-mediated organometallic catalysis through electronically excited nickel(II).” Welin, E. R.; Le, C.; Lin, S.; Arias-Rotondo, D. M.; McCusker, J. K.; MacMillan, D. W. C. Science 2017, 355, 380.
“Selective sp3 C–H Alkylation via Polarity Match Based Cross-Coupling.” Le, C.; Liang, Y.; Evans, R. W.; Li, X.; MacMillan, D. W. C. Nature 2017, 547, 79.
“Native functionality in triple catalytic cross-coupling: sp3 C–H bonds as latent nucleophiles.” Shaw, M. S.; Shurtleff, V. W.; Terrett, J. A.; Cuthbertson, J. D.; MacMillan, D. W. C. Science 2016, 352, 1304.
“Aryl amination using ligand-free Ni(II) salts and photoredox catalysis.” Corcoran, E. B.; Pirnot, M. T.; Lin, S.; Dreher, S. D.; DiRocco, D. A.; Davies, I. W.; Buchwald, S. L.; MacMillan, D. W. C. Science 2016, 353, 279.
“Metallaphotoredox-catalyzed sp3–sp3 cross-coupling of carboxylic acids with alkyl halides” Johnson, C. P.; Smith, R. T.; Allmendinger, S.; MacMillan, D. W. C. Nature 2016, 536, 322.
“Switching on Elusive Organometallic Mechanisms with Photoredox Catalysis.” Terrett, J. A.; Cuthbertson, J. D.; Shurtleff, V. W.; MacMillan, D. W. C. Nature 2015, 524, 330.
“Alcohols as alkylating agents in heteroarene C–H functionalization.” Jin, J.; MacMillan, D. W. C. Nature 2015, 525, 87.
“Merging photoredox with nickel catalysis: Coupling of a-carboxyl sp3-carbons with aryl halides.” Zuo, Z.; Ahneman, D.; Chu, L.; Terrett, J.; Doyle, A. G.; MacMillan, D. W. C. Science 2014, 345, 437.
“Trifluoromethylation of Arenes and Heteroarenes via Photoredox Catalysis.” Nagib, D. A.; MacMillan, D. W. C. Nature 2011, 480, 224.
“Enantioselective a-Trifluoromethylation of Aldehydes via Photoredox Organocatalysis.” Nagib, D. A.; Scott, M. E.; MacMillan, D. W. C. Am. Chem. Soc. 2009, 131, 10875.
“Merging Photoredox Catalysis with Organocatalysis: The Direct Asymmetric Alkylation of Aldehydes.” Nicewicz, D. A.; MacMillan, D. W. C. Science 2008, 322, 77.
“Enantioselective Organocatalytic Hydride Reduction.” Ouellet, S. G.; Tuttle, J. B.; MacMillan, D. W. C. Am. Chem. Soc. 2005, 127, 32.
“Two-Step Synthesis of Carbohydrates by Selective Aldol Reactions.” Northrup, A. B.; MacMillan, D. W. C. Science 2004, 305, 1752.
“Enantioselective Organocatalytic Indole Alkylations. Design of a New and Highly Effective Chiral Amine for Iminium Catalysis.” Austin, J. A.; MacMillan, D. W. C. Am. Chem. Soc. 2002, 124, 1172.
“The First Direct and Enantioselective Cross-Aldol Reaction of Aldehydes.” Northrup, A. B.; MacMillan, D. W. C. Am. Chem. Soc. 2002, 124, 6798.
“New Strategies for Organic Catalysis: The First Highly Enantioselective Organocatalytic Diels–Alder Reaction.” Ahrendt, K. A.; Borths, C. J.; MacMillan, D. W. C. Am. Chem. Soc. 2000, 122, 4243.
Related News

The Rabinowitz Lab: In Partnership with the Ludwig Institute for Cancer Research

MacMillan Lab’s couple-close advance “makes the impossible, possible”


